I’ve spent the evening knocking out this little sketch to scrape molecular information from ChemSpider to inform my mass spectrometry. Until now I’ve been doing this by hand or getting the students to, when they are able, but this is the start of my attempt to automate the process. Ultimately I will just want to feed it a text file of compound names and it will parse the output into a document.
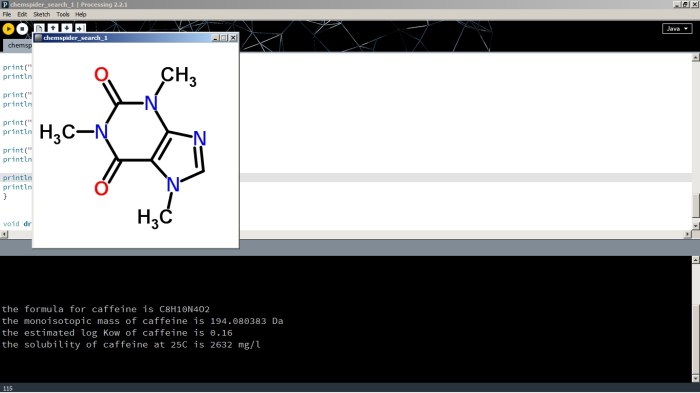
Open a Processing window, copy the code below into it, edit the first string to contain the name of the compound of your choice and click run. Here’s an image of example output.
// sketch to grab molecular information from a named compound from ChemSpider
// AUT School of Applied Science
// June 2015
// drchrispook@gmail.com
// released under GPLv3 licence – https://gnu.org/licenses/quick-guide-gplv3.html// ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~
// enter the compound name you want information for
String compound = “caffeine”;
// ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~ ~// sketch continues
PImage webImg;String URL = “http://www.chemspider.com//Search.aspx?q=”;
String nameURL = URL + compound;
String[] html1 = loadStrings(nameURL);String CSID = “CSID”;
String formula = “formula”;
String MIM = “MIM”;
String solubility = “solubility”;
String logKow = “logKow”;
int imageSize = 400;
void setup() {
size(imageSize, imageSize);for(String i:html1) { // iterate through each string in the array
// println(i);
int line = i.indexOf(“CSID:”); // identifies CSID line by searching for this stringif(line > 0) { // if you find that string
String[] trim1 = split(i, ‘,’); // split off the junk
// for(int i = 0; i < trim1.length; i++) { // println(trim[i]); // } String[] trim2 = split(trim1[0], ‘:’); // split off more junk CSID = trim2[1]; // set the CSID break; } } String imageURL = “http://www.chemspider.com//ImagesHandler.ashx?id=” + CSID + “&w=” + imageSize + “&h=” + imageSize; webImg = loadImage(imageURL, “jpg”); String CSIDURL = “http://www.chemspider.com//Chemical-Structure.” + CSID + “.html”; // use the CSID to construct the url to that molecule’s entry String[] html2 = loadStrings(CSIDURL); // grab the html //println(html2); for(String i:html2) { // this is the main part of the macro // println(i); int MIMLine = i.indexOf(“Monoisotopic mass”); // identifies MIM line int formulaLine = i.indexOf(“Molecular-Formula”); // identifies the formula line int solubilityLine = i.indexOf(“Solubility at 25 deg C (mg/L):”); // etc int logKowLine = i.indexOf(“Log Kow (KOWWIN”); // etc if(formulaLine > 0) {
String[] trim1 = split(i, ‘/’);
// for(int e = 0; e < trim1.length; e++) { // println(trim1[e]); // } String[] trim2 = split(trim1[2], ‘”‘); formula = trim2[0]; } if(MIMLine > 0) {
String[] trim1 = split(i, ‘>’);
// for(int e = 0; e < trim1.length; e++) { // println(trim1[e]); // } String[] trim2 = split(trim1[3], ‘ ‘); MIM = trim2[0]; } if(solubilityLine > 0) {
String[] trim1 = split(i, ‘ ‘);
// for(int e = 0; e < trim1.length; e++) { // println(trim1[e]); // } solubility = trim1[12]; } if(logKowLine > 0) {
String[] trim1 = split(i, ‘ ‘);
// for(int e = 0; e < trim1.length; e++) {
// println(trim1[e]);
// }
logKow = trim1[11];
}
}
print(“the formula for ” + compound + ” is “);
println(formula);print(“the monoisotopic mass of ” + compound + ” is “);
println(MIM + ” Da”);print(“the estimated log Kow of ” + compound + ” is “);
println(logKow);print(“the solubility of ” + compound + ” at 25C is “);
println(solubility + ” mg/l”);
}
void draw() {
background(0);
image(webImg, 0, 0);
}